GPR-GNN
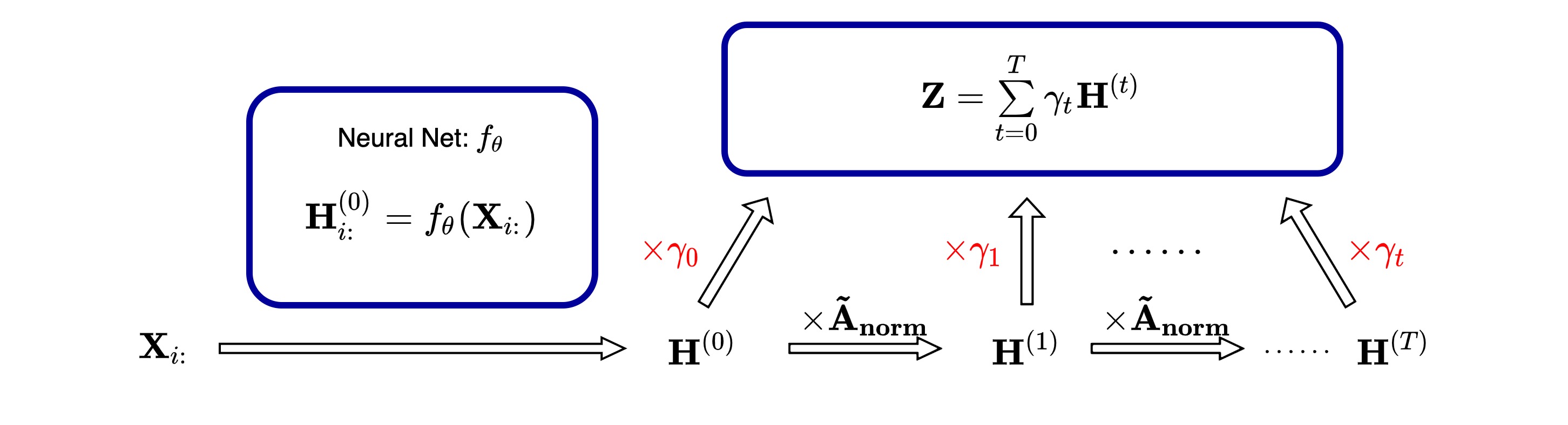
Structure
Results
Run with following (available dataset: "cora", "citeseer", "pubmed", "computers", "photo", "squirrel", "chameleon", "cornell", "texas")
sh Reproduce_GPRGNN.sh
|
Cora |
Citeseer |
PubMed |
Computers |
Photo |
Chameleon |
Squirrel |
Texas |
Cornell |
| Tensorflow |
79.08 |
65.96 |
83.74 |
83.44 |
92.06 |
67.61 |
51.82 |
88.02 |
85.25 |
| Paddle |
|
|
|
|
|
|
|
|
|
| Origin |
79.51 |
67.63 |
85.07 |
82.90 |
91.93 |
67.48 |
49.93 |
92.92 |
91.36 |
Details
GPR-GNN



Convolutional layer class
GPRConv
In file layers/conv/gpr_conv.py
The basic structure is similar to gcn_conv.py
- Provides the GPR-GNN conv layer
- In
__init__ method: Add the initialization for learnt weights $\gamma_k$ and define the steps of propagation $K$
- For initializaiton method provided:
- $\textbf{SGC}$: $\gamma_k = \delta_{kK}$, weight for the last layer is set to $1$, others as $0$;
- $\textbf{PPR}$: $\gamma_k=\alpha(1-\alpha)^{k}\text{ for } k<K \text{ and }\gamma_K=(1-\alpha)^K$;
- $\textbf{NPPR}$: $\gamma_k=\alpha^{k} / \sum\limits_{k=0}^{K}\alpha^{k}$;
- $\textbf{Random}$: $\gamma\sim 2\sqrt{\frac{K+1}{3}}\mathbf{U}(-\sqrt{\frac{3}{K+1}},\sqrt{\frac{3}{K+1}})$
- $\textbf{WS}$: User defined initial value for $\gamma_k$
- For $K$: recommended value is 10 (proposed by GPR-GNN origin paper)
- In
reset_parameters method: Use $\textbf{PPR}$ initializaiton method to reset parameters
Model class
GPRGNNModel
In file models/gprgnn.py
- Provides the complete GPR-GNN model
- In
__init__ method: besides basic dimension information of node features and num of classes, GPR related hyperparameters ($K$, $\textbf{Init}$, $\alpha$, $\text{dprate}$, etc.) are also transfered to the network
- In
forward propagation:
- After feature extraction, it passes a special dropout layer controled by $\text{dprate}$ before entering propagation layers.
- Then, those representations of nodes are transferred into
GPRConv
Transforms class
NormalizeFeatures
In file transforms/normalize_features.py
- Provides a row-wise normalization for feature matrix
Trainer
In file examples/gprgnn/gprgnn_trainer.py
- Function
random_planetoid_splits is defined to randomly split dataset and ensure the numbers of nodes from each classes are the same in training set.
- Maximum of epoches is set as 1000 and early stopping trick is applied.
- Early stopping mechanism is used.
Experiment
In file examples/gprgnn/Reproduce_GPRGNN.sh
-
To keep the same with experiments in origin paper,5 homophily graph datasets, Cora, Citeseer, PubMed, Computers and Photo , adopt sparse split $(2.5%/2.5%/95%)$ for (training/validation/test), and 4 heterophily graph datasets, Chameleon, Squirrel, Texas and Cornell, adopt dense split $(60%/20%/20%)$ .
-
Train model with each hyperparameter setting for 10 times and calculate the average accuracy.
-
Best hyperparameters for different datasets are given in the file.
Result
In file examples/gprgnn/test.accuracy
|
Cora |
Citeseer |
PubMed |
Computers |
Photo |
Chameleon |
Squirrel |
Texas |
Cornell |
| Tensorflow |
79.08 |
65.96 |
83.74 |
83.44 |
92.06 |
67.61 |
51.82 |
88.02 |
85.25 |
| Paddle |
|
|
|
|
|
|
|
|
|
| Origin |
79.51 |
67.63 |
85.07 |
82.90 |
91.93 |
67.48 |
49.93 |
92.92 |
91.36 |
- Test accuracy on
Texas and Cornell is unstable, fluctuating from 75 ~ 95.

