




Note: BrainPy is a project under development. More features are coming soon. Contributions are welcome.
Why to use BrainPy
BrainPy is an integrative framework for computational neuroscience and brain-inspired computation. Three core functions are provided in BrainPy:
- General numerical solvers for ODEs and SDEs (future will support DDEs and FDEs).
- Neurodynamics simulation tools for brain objects, such like neurons, synapses and networks (future will support soma and dendrites).
- Neurodynamics analysis tools for differential equations, including phase plane analysis and bifurcation analysis (future will support continuation analysis and sensitive analysis).
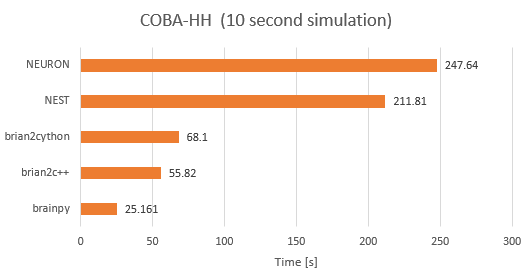
Moreover, BrainPy can effectively satisfy your basic requirements: 1. Easy to learn and use, because it is only based on Python language and has little dependency requirements; 2. Highly flexible and transparent, because it endows the users with the fully data/logic flow control; 3. Simulation can be guided with the analysis, because the same code in BrainPy can not only be used for simulation, but also for dynamics analysis; 4. Efficient running speed, because BrainPy is compatitable with the latest JIT compilers (or any other computing backend you prefer).
Installation
Install BrainPy by using pip:
> pip install --pre brainpy-simulator
Install BrainPy by using conda:
> conda install brainpy-simulator -c brainpy
Install BrainPy from source:
> pip install git+https://github.com/PKU-NIP-Lab/BrainPy
> # or
> pip install git+https://git.openi.org.cn/OpenI/BrainPy
> # or
> pip install -e git://github.com/PKU-NIP-Lab/BrainPy.git@V0.2.5
BrainPy is based on Python (>=3.7), and the following packages are required to be installed to use BrainPy:
- NumPy >= 1.13
- Matplotlib >= 3.2
Neurodynamics simulation

|
The Hodgkin–Huxley model, or conductance-based model,
is a mathematical model that describes how action potentials
in neurons are initiated and propagated. It is a set of nonlinear
differential equations that approximates the electrical characteristics
of excitable cells such as neurons and cardiac myocytes.
|
|
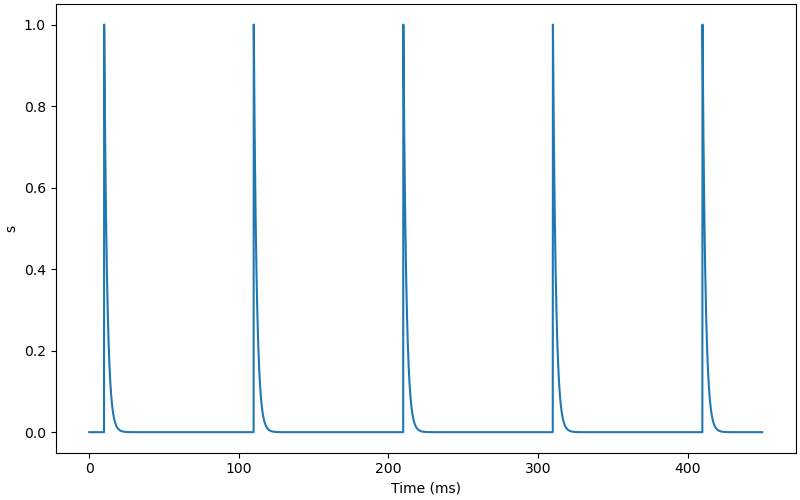
AMPA synapse model.
|
|
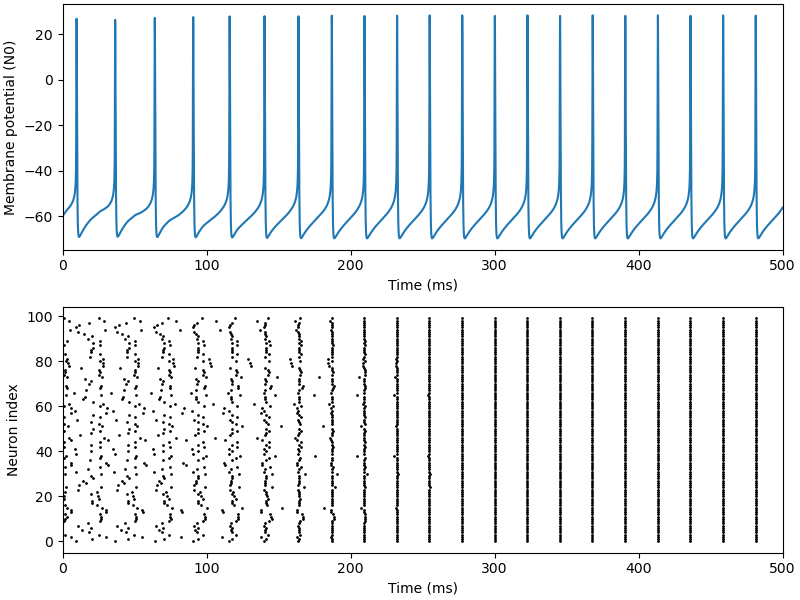
Implementation of the paper: Wang, Xiao-Jing, and György Buzsáki. “Gamma oscillation by
synaptic inhibition in a hippocampal interneuronal network
model.” Journal of neuroscience 16.20 (1996): 6402-6413.
|
|
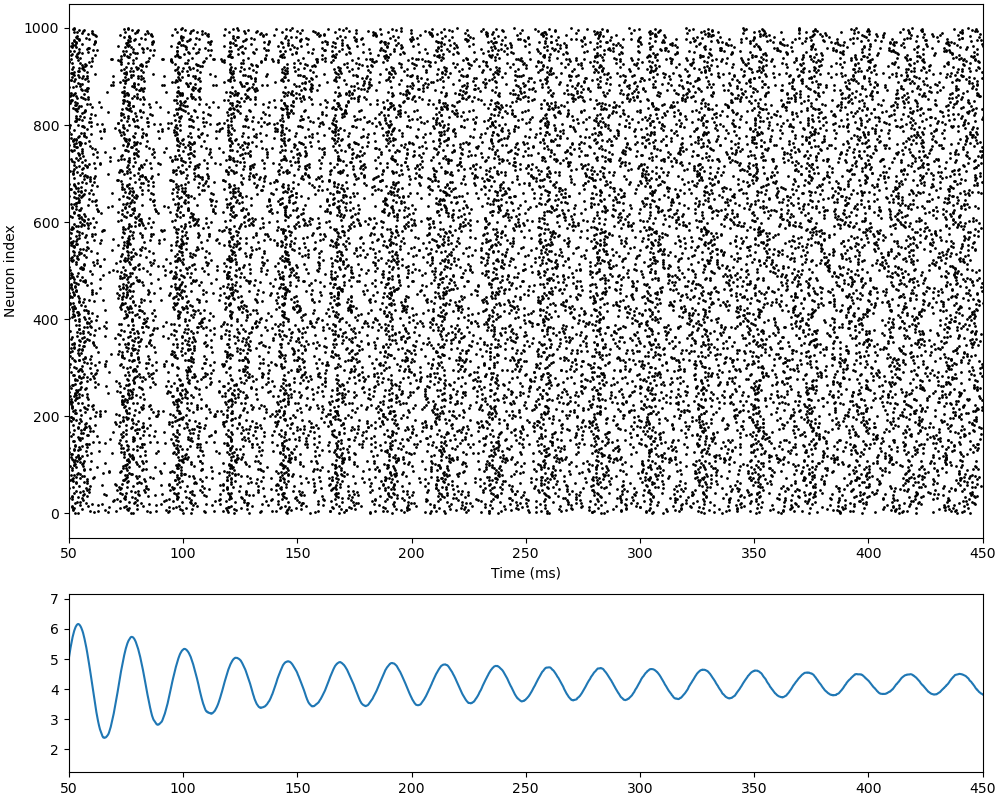
Implementation of the paper: Van Vreeswijk, Carl, and Haim Sompolinsky.
“Chaos in neuronal networks with balanced excitatory and inhibitory activity.”
Science 274.5293 (1996): 1724-1726.
|
|
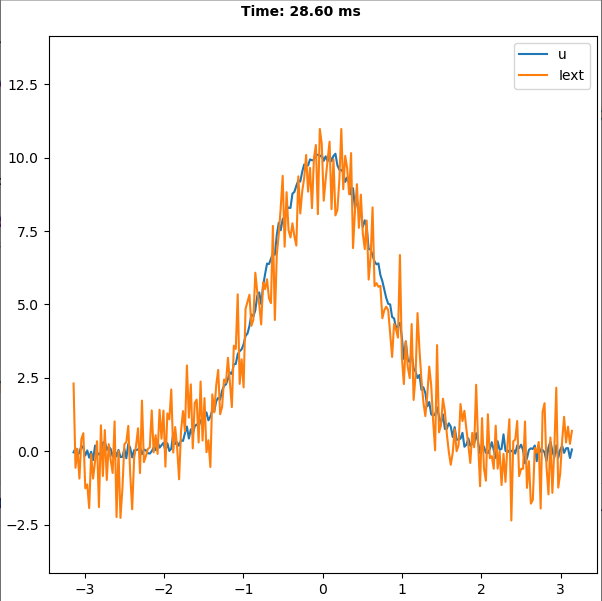
Implementation of the paper: Si Wu, Kosuke Hamaguchi, and Shun-ichi Amari. "Dynamics and
computation of continuous attractors." Neural
computation 20.4 (2008): 994-1025.
|
More neuron examples please see BrainPy-Models/neurons;
More synapse examples please see BrainPy-Models/synapses;
More network examples please see BrainPy-Models/from_papers.
Neurodynamics analysis
|
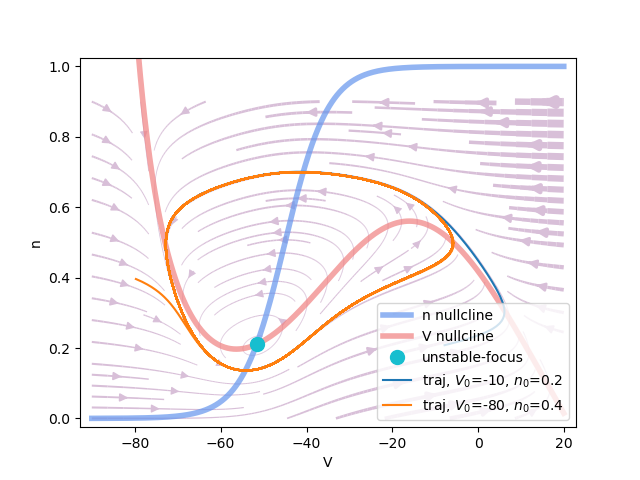
Phase plane analysis of the INa,p+-IK model, where
"input" is 50., and "Vn_half" is -45..
|
|
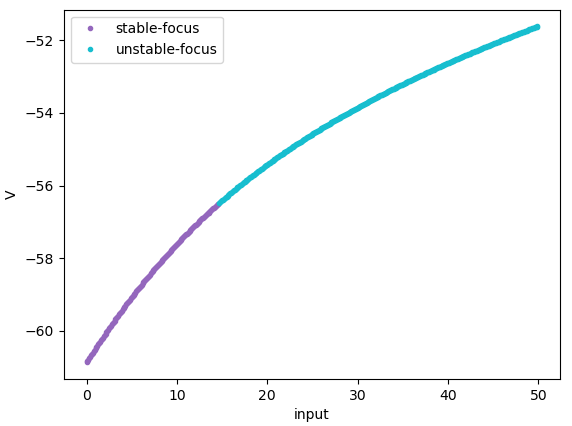
Codimension 1 bifurcation analysis of the INa,p+-IK model,
in which "input" is varied in [0., 50.].
|
|
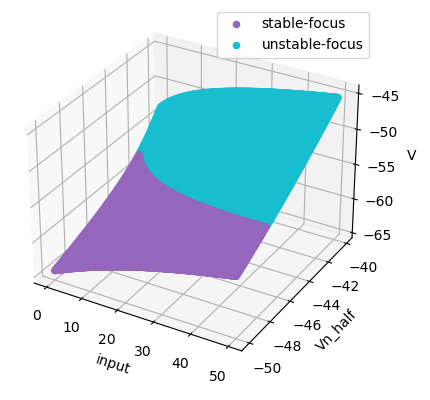
Codimension 2 bifurcation analysis of a two-variable neuron model:
the INa,p+-IK model, in which "input" is varied
in [0., 50.], and "Vn_half" is varied in [-50, -40].
|
|
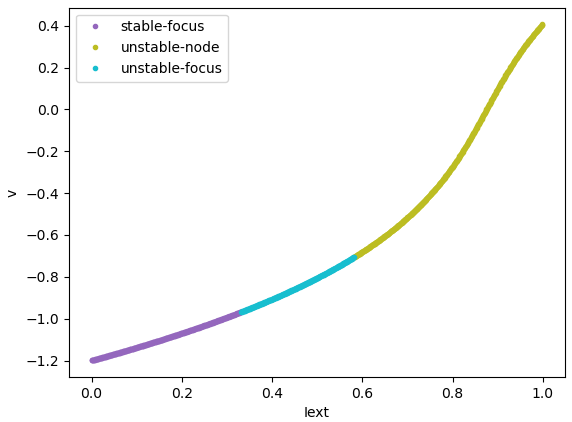
Codimension 1 bifurcation analysis of FitzHugh Nagumo model, in which
"a" is equal to 0.7, and "Iext" is varied in [0., 1.].
|
|
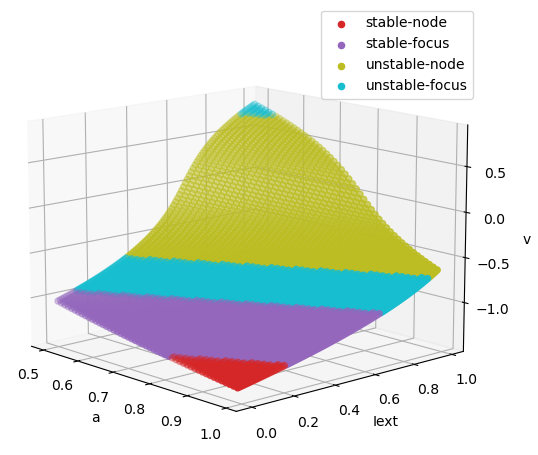
Codimension 2 bifurcation analysis of FitzHugh Nagumo model, in which "a"
is varied in [0.5, 1.0], and "Iext" is varied in [0., 1.].
|
More examples please see BrainPy-Models/dynamics_analysis.












