Are you sure you want to delete this task? Once this task is deleted, it cannot be recovered.
|
|
2 months ago | |
|---|---|---|
| .. | ||
| cmake | 3 months ago | |
| docs | 3 months ago | |
| sciai | 2 months ago | |
| tutorial | 3 months ago | |
| .env | 3 months ago | |
| CMakeLists.txt | 3 months ago | |
| LICENSE | 3 months ago | |
| NOTICE | 3 months ago | |
| README.md | 3 months ago | |
| README_CN.md | 3 months ago | |
| RELEASE.md | 3 months ago | |
| RELEASE_CN.md | 3 months ago | |
| build.sh | 3 months ago | |
| install.sh | 3 months ago | |
| requirements.txt | 3 months ago | |
| setup.py | 3 months ago | |
| version.txt | 3 months ago | |
README.md
ENGLISH | 简体中文
MindSpore SciAI
- MindSpore SciAI Introduction
- Model Library
- Installation Guide
- Quick Start
- Documents
- Community
- Contribution
- Release Notes
- License
MindSpore SciAI Introduction
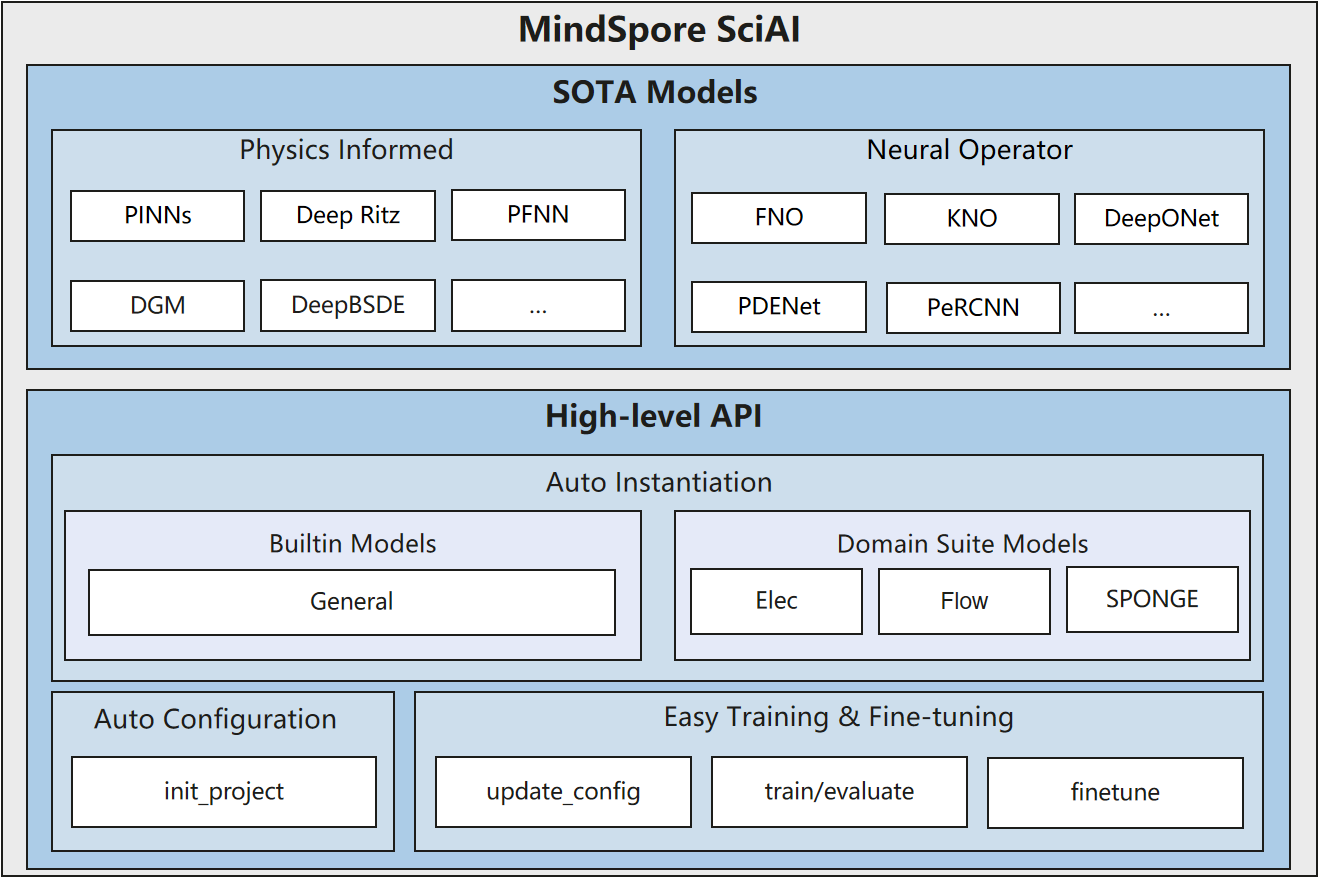
Based on MindSpore, SciAI is a model library with 60 built-in most frequently used and cited AI4Sci(AI for Science) models, which cover SOTA models from physics-informed (PINNs, DeepRitz, PFNN, etc.) to neural operators (FNO, DeepONet, PDENet), ranking No.1 in the world in terms of coverage.
MindSpore SciAI provides the developers and users with a high-level API (auto configuration, auto instantiation, training and fine-tuning, etc.), allowing an immediate deployment.
With these features, MindSpore SciAI covers a wide variety of scientific computation fields,
including fluid dynamics, electromagnetism, sound, heat, solid and biology, providing developers and users with an efficient and convenient AI4SCI computation platform.
The hierarchy of project SciAI is given below.
├── SciAI
│ ├── cmake # compilation related
│ ├── docs # documentation
│ │ ├── FAQ # frequently asked questions
│ ├── sciai # SciAI framework
│ │ ├── architecture # base neural networks modules
│ │ ├── common # common modules
│ │ ├── context # context functions
│ │ ├── model # **AI4SCI high-frequency model library**
│ │ ├── operators # high order derivatives
│ │ └── utils # Other auxiliary functions
│ └── tutorial # tutorial models
Model Library
SciAI model library provides a wide variety of models that are frequently used and cited in scientific computation.
The following table summarizes the current available neural networks and their corresponding domains.
| Domain | Network | MindSpore Implementation and Parameters | Ascend | GPU |
|---|---|---|---|---|
| General Physics | auq_pinns | link | ✅ | ✅ |
| General Physics | cpinns | link | ✅ | ✅ |
| General Physics | deep_hpms | link | ✅ | ✅ |
| General Physics | deep_ritz | link | ✅ | ✅ |
| General Physics | deepbsde | link | ✅ | |
| General Physics | deeponet | link | ✅ | ✅ |
| General Physics | dgm | link | ✅ | ✅ |
| General Physics | fbsnns | link | ✅ | ✅ |
| General Physics | fpinns | link | ✅ | ✅ |
| General Physics | gradient_pathologies_pinns | link | ✅ | ✅ |
| General Physics | hp_vpinns | link | ✅ | ✅ |
| General Physics | laaf | link | ✅ | ✅ |
| General Physics | mgnet | link | ✅ | ✅ |
| General Physics | multiscale_pinns | link | ✅ | ✅ |
| General Physics | pfnn | link | ✅ | |
| General Physics | phygeonet | link | ✅ | ✅ |
| General Physics | pi_deeponet | link | ✅ | |
| General Physics | pinns | link | ✅ | |
| General Physics | pinns_ntk | link | ✅ | ✅ |
| General Physics | ppinns | link | ✅ | ✅ |
| General Physics | xpinns | link | ✅ | ✅ |
| Hamiltonian Systems | sympnets | link | ✅ | ✅ |
| Fluid Dynamic | hfm | link | ✅ | ✅ |
| Fluid Dynamic | label_free_dnn_surrogate | link | ✅ | ✅ |
| Fluid Dynamic | nsf_nets | link | ✅ | ✅ |
| Fluid Dynamic | *burgers_fno | link | ✅ | ✅ |
| Fluid Dynamic | *burgers_kno | link | ✅ | ✅ |
| Fluid Dynamic | *navier_stokes_fno | link | ✅ | ✅ |
| Fluid Dynamic | *navier_stokes_kno | link | ✅ | ✅ |
| Fluid Dynamic | *navier_stokes_3d_fno | link | ✅ | ✅ |
| Fluid Dynamic | *pde_net | link | ✅ | ✅ |
| Fluid Dynamic | *percnn | link | ✅ | ✅ |
| Elastodynamics | pinn_elastodynamics | link | ✅ | ✅ |
| Thermodynamics | pinn_heattransfer | link | ✅ | ✅ |
| Meteorology | enso | link | ✅ | ✅ |
| Geology | inversion_net | link | ✅ | ✅ |
| Geology | pinn_helmholtz | link | ✅ | ✅ |
| Oceanic Physics | ocean_model | link | ✅ | |
| Oceanic Physics | pinns_swe | link | ✅ | ✅ |
| Electromagnetism | maxwell_net | link | ✅ | ✅ |
| Electromagnetism | *AD_FDTD_invert_f | link | ✅ | |
| Electromagnetism | *AD_FDTD_microstrip_filter | link | ✅ | |
| Electromagnetism | *AD_FDTD_inverse | link | ✅ | |
| Electromagnetism | *frequency_domain_maxwell | link | ✅ | ✅ |
| Electromagnetism | *frequency_domain_maxwell_3D_dielectric_slab | link | ✅ | ✅ |
| Electromagnetism | *frequency_domain_maxwell_3D_waveguide_cavity | link | ✅ | ✅ |
| Electromagnetism | *meta_auto_decoder | link | ✅ | ✅ |
| Electromagnetism | *pinn_fwi | link | ✅ | ✅ |
| Electromagnetism | *SED_ANN | link | ✅ | ✅ |
| Electromagnetism | *time_domain_maxwell | link | ✅ | ✅ |
| Electromagnetism | *metasurface_holograms | link | ✅ | ✅ |
| Biology | *MEGA-Fold | link (inference) link (training) | ✅ | ✅ |
| Biology | *MEGA-EvoGen | link | ✅ | ✅ |
| Biology | *MEGA-Assessment | link (inference) link (training) | ✅ | ✅ |
| Biology | *ColabDesign | link | ✅ | ✅ |
| Biology | *DeepFRI | link | ✅ | ✅ |
| Biology | *Multimer | link | ✅ | ✅ |
| Biology | *ProteinMPNN | link | ✅ | ✅ |
| Biology | *UFold | link | ✅ | ✅ |
| Biology | *esm-if1 | link | ✅ | ✅ |
| Biology | *esm2 | link | ✅ | ✅ |
| Biology | *grover | link | ✅ | ✅ |
Note: the "*" in the model names indicates that these models have already been released at an earlier time by MindSpore
and MindScience.
Installation Guide
Confirming the System Environment Information
Due to the dependency between SciAI and MindSpore, please follow the table below and install the corresponding MindSpore
version from MindSpore Download Guide.
| SciAI Version | Branch | MindSpore Version | Python Version |
|---|---|---|---|
| 0.1.0 | master | >=2.2.0 | =3.7 |
For other dependencies, see requirements.txt,
and they can be installed via the following command.
pip install -r requirements.txt
The supported hardware and operating systems are listed below.
| Hardware | OS | Status |
|---|---|---|
| Ascend 910 | Ubuntu-x86 | ✅ |
| Ubuntu-aarch64 | ✅ | |
| EulerOS-aarch64 | ✅ | |
| CentOS-x86 | ✅ | |
| CentOS-aarch64 | ✅ | |
| GPU CUDA 11.1 | Ubuntu-x86 | ✅ |
Method 1: Install With Pip
This method installs SciAI from .whl package automatically downloaded from MindSpore website,
which does not require the download and compilation of source code.
Note: This installation method is not currently supported.
It will be supported after Mindspore 2.2.0 official release.
pip install https://ms-release.obs.cn-north-4.myhuaweicloud.com/2.2.0/MindScience/sciai/gpu/{arch}/cuda-11.1/sciai-{version}-cp37-cp37m-linux_{arch}.whl -i https://pypi.tuna.tsinghua.edu.cn/simple
- When the network is connected, dependencies of the SciAI installation package are automatically downloaded during
the .whl package installation. For details about dependencies, see setup.py.- {version} denotes the version of SciAI. For example, when you are installing SciAI 0.1.0, {version} should
be0.1.0.- {arch} denotes the system architecture. For example, the Linux system you are using is x86 architecture 64-bit,
{arch} should bex86_64. If the system is ARM architecture 64-bit, then it should beaarch64.
The following table provides the corresponding installation commands to each architecture and Python version.
| Device | Architecture | Python | Command |
|---|---|---|---|
| Ascend | x86_64 | Python=3.7 | pip install https://ms-release.obs.cn-north-4.myhuaweicloud.com/2.2.0/MindScience/sciai/gpu/x86_64/cuda-11.1/sciai-0.1.0-cp37-cp37m-linux_x86_64.whl -i https://pypi.tuna.tsinghua.edu.cn/simple |
| aarch64 | Python=3.7 | pip install https://ms-release.obs.cn-north-4.myhuaweicloud.com/2.2.0/MindScience/sciai/ascend/aarch64/sciai-0.1.0-cp37-cp37m-linux_aarch64.whl -i https://pypi.tuna.tsinghua.edu.cn/simple |
|
| GPU | x86_64 | Python=3.7 | pip install https://ms-release.obs.cn-north-4.myhuaweicloud.com/2.2.0/MindScience/sciai/gpu/x86_64/cuda-11.1/sciai-0.1.0-cp37-cp37m-linux_x86_64.whl -i https://pypi.tuna.tsinghua.edu.cn/simple |
Note: If you have other MindScience package(s) installed in your conda or python env, such as MindElec, MindFlow
, MindSponge,
please uninstall the MindScience package(s) in the environment first to avoid pip behavior conflicts.
Method 2: Install From Source Code
-
Clone the source code from the Git repository of MindScience.
cd ~ git clone https://gitee.com/mindspore/mindscience.git -
Build SciAI with script
build.sh.cd mindscience/SciAI bash build.sh -j8 -
Install the
.whlpackagebash install.sh
Installation Verification
To verify the installation, run the following commands. If the error message No module named 'sciai' is not displayed,
the installation is successful.
python -c 'import sciai'
Quick Start
The model library SciAI provides two means of model training and evaluation,
providing the developers with flexible and simple usage and development interface.
Method 1: Launch by Advanced API (recommended)
After installation of SciAI:
- Train and Finetune with AutoModel
Users can call the function AutoModel.from_pretrained to get the network models, which are supported in SciAI.
Users can use the function AutoModel.train to train the neural networks, and before training,
users can use AutoModel.update_config to configure the training parameters
or finetune the model by loading the .ckpt file.
The acceptable arguments for AutoModel.update_config depend on the model instantiated.
User can refer to the links Model Library,
column MindSpore Implementation and Parameters for detailed information.
from sciai.model import AutoModel
# obtain the `cpinns` model.
model = AutoModel.from_pretrained("cpinns")
# train the network with default configuration.
# the figures, data and logs generated will be saved in your execution path.
model.train()
# or load a `.ckpt` file.
model.update_config(load_ckpt=True, load_ckpt_path="./checkpoints/your_checkpoint_file.ckpt", epochs=500)
# continue training the model based on the loaded parameters.
model.train()
- Evaluate with AutoModel
Users can evaluate the trained/pre-trained networks with the function AutoModel.evaluate.
This function will load the .ckpt files provided in SciAI by default.
Alternatively, user can load their own .ckpt file with the interface AutoModel.update_config.
from sciai.model import AutoModel
# obtain the `cpinns` model
model = AutoModel.from_pretrained("cpinns")
# load the default ckpt file and evaluate the model
model.evaluate()
# or load the ckpt file provided by the user(Optional)
model.update_config(load_ckpt=True, load_ckpt_path="./checkpoints/your_checkpoint_file.ckpt")
# and then evaluate the model
model.evaluate()
- View training and validation results
Users can find a directory named auto_model_xxx in the script running directory, which contains checkpoints, data
, figures, logs four subdirectories, containing checkpoints, data, pictures, and logs respectively.
Users can view training results in these directories. For example:
├── auto_model_cpinns
│ ├── checkpoints # checkpoint files
│ ├── data # data files
│ ├── figures # plot figures
│ └── logs # log files
- 可配置参数:
AutoModel.update_config(**kwargs) 支持修改网络参数,包括学习率、训练周期、混合精度等级、动静态图、数据读取保存路径、checkpoints文件加载路径等,
具体请参考网络模型库中MindSpore实现与网络参数链接。常用的可配置参数见下表:
- Configurable parameters:
AutoModel.update_config(**kwargs) supports modifying network parameters, including learning rate, training epochs,
auto mixed precision level, graph or pynative mode, data loading and saving path, checkpoints file loading path, etc.
For details, please refer to the link in MindSpore implementation and network parameters column
in Model Library. Commonly used configurable parameters are shown in the table below:
| parameter | description | default values |
|---|---|---|
| save_ckpt | whether save checkpoint or not | true |
| save_fig | whether save and plot figures or not | true |
| load_ckpt | whether load checkpoint or not | false |
| save_ckpt_path | checkpoint saving path | ./checkpoints |
| load_ckpt_path | checkpoint loading path | ./checkpoints/model_final.ckpt |
| load_data_path | path to load data | ./data |
| save_data_path | path to save data | ./data |
| figures_path | figures saving path | ./figures |
| log_path | log saving path | ./logs |
| print_interval | time and loss print interval | 10 |
| ckpt_interval | checkpoint saving interval | 1000 |
| lr | learning rate | 8e-4 |
| epochs | number of epochs | 15001 |
| download_data | necessary dataset and/or checkpoints | cpinns |
| force_download | whether download the dataset or not by force | false |
| amp_level | MindSpore auto mixed precision level | O3 |
| device_id | device id to set | None |
| mode | MindSpore Graph mode(0) or Pynative mode(1) | 0 |
- Recover Checkpoints and Dataset
Users may encounter the situation that the automatically downloaded checkpoints or dataset are missing or deleted,
while they can always be re-downloaded by updating the config as follows.
model.update_config(force_download=True)
Method 2: Launch by Source code
Users can clone the whole repository and prepare the environment with the following commands.
Running .env will append the path of SciAI directory to the environment variable PYTHONPATH.
git clone https://gitee.com/mindspore/mindscience
source ./mindscience/SciAI/.env
Then train or evaluate a model in Model Library according to the Quick Start in README.md of the
model. Let's take model cpinns as an example:
- Train the Model:
Users can run script train.py to train the model.
cd ./mindscience/SciAI/sciai/model/cpinns/
python ./train.py [--parameters]
- Finetune the Model with
.ckptfile:
python ./train.py --load_ckpt true --load_ckpt_path {your_file}.ckpt [--parameters]
- Evaluate the Model:
Users can run script eval.py to evaluate the trained model. Take cpinns as an example:
python ./eval.py [--parameters]
Configurable parameters [--parameters] include the learning rate, training epochs, data saving and loading paths,
checkpoint loading path and so on.
For details, see the links provided in the MindSpore Implementation and Parameters column
in Model Library.
- Recover Checkpoints and Dataset
Users may encounter with the situation that the automatically downloaded checkpoints or dataset are missing or deleted,
while they can always be re-downloaded with the following solutions:
- Edit the configuration file
config.yamlin the model directory, and set theforce_downloadto true. - Append the extra parameter
--force_download Truewhen launchingtrain.pyoreval.pyin the terminal.
Documents
For more details about the installation guides, tutorials, and APIs,
see SciAI Documents.
If you encounter problems with SciAI, please refer to FAQ.
Community
Contribution
Make your contribution. For more details, please refer to
out Contributor Wiki
Release Notes
License
MindScience is scientific computing kits for various industries based on the converged MindSpore framework.
Jupyter Notebook Python Unity3D Asset Pickle nesC other
